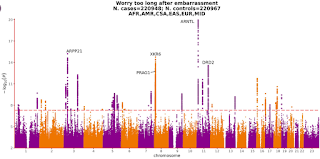
As I have discussed in a previous blog post, despite claims of strong heritability from twin studies and tepid results from genetic studies to date, there has been little success in elucidating causal genetic architecture for schizophrenia. A recent article comments on this issue (30 years too late).
The Human Genome Project was undertaken primarily to discover genetic causes and better treatments for human diseases. Schizophrenia was targeted since three of the project`s principal architects had a personal interest and also because, based on family, adoption, and twin studies, schizophrenia was widely believed to be a genetic disorder. Extensive studies using linkage analysis, candidate genes, genome wide association studies [GWAS], copy number variants, exome sequencing and other approaches have failed to identify causal genes. Instead, they identified almost 300 single nucleotide polymorphisms [SNPs] associated with altered risks of developing schizophrenia as well as some rare variants associated with increased risk in a small number of individuals. Risk genes play a role in the clinical expression of most diseases but do not cause the disease in the absence of other factors. Increasingly, observers question whether schizophrenia is strictly a genetic disorder.
An argument that is often made to counter this three decade failure, is that schizophrenia might be caused by rare genetic variants that, to date, have not been identified in genetic studies, but might be in the future. Bolstering this premise is the fact that individuals with rare genetic disorders such as 22q11.2 Deletion Syndrome and Fragile X Syndrome have a much higher prevalence for the diagnosis of schizophrenia. Such genetic disorders have been fairly easy to identify and most of the individuals have other issues such as intellectual disabilities and seizures. These disorders generally involve larger portions of DNA than a single genetic variant, however, so it is also difficult to identify which genes might be responsible.
Along with being a schizophrenia risk gene, CACNA1G is also associated with the risk of severe intellectual or developmental disability.
In fact, all of the rare variants noted in the paper also confer risk for developmental and intellectual disabilities. If one had a gene that only conferred a risk for schizophrenia and nothing else, the case would be stronger. However, the fact that these patients generally have other developmental and intellectual disablities is not a coincidence. It speaks to the reality of clinical psychiatry and needs to be looked at in a broader social context.
Patients with intellectual disabilities are often put into the mental health system due to behavioral issues and other coping difficulties that anyone has witnessed. These issues will get them into difficulties both in the home with those involved in their care and other social environments. The hope is that some intervention, generally medications, will help keep them under control and out of troubles that might affect their ability to find basic living arrangements, hold jobs, etc. As a psychiatrist, there is no clinical justification for just keeping someone medicated without an appropriate diagnosis in the Diagnostic and Statistical Manual (DSM). The number of diagnoses one can give that would justify the kind of medications being considered in such a scenario, which would be sedative antipsychotic medications like Haldol or Olanzapine, or mood stabilizers like lithium and Depakote are few.
So we have a situation where the questions will be structured (consciously or unconsciously) to achieve the goal. “Billy, did the voices tell you to break the window?,’ “Jane, do you feel like people are making fun of you?,” “Mike just had a mood outburst,” etc. Soon you have a patient who hears voices, is paranoid and has mood issues. Then you give a diagnosis of schizophrenia, or more often, schizoaffective disorder. The fact of the matter is that these “symptoms” are simply not the same as someone with the classic diagnosis of schizophrenia, who hears actual voices speaking to him as opposed to some impulse, who has a paranoid conspiracy brewing in his head related to the CIA or the Masons or the like rather than the very real concern of someone with a mental disability that people are making fun of them. Likewise, someone with classic manic symptoms where they are awake for a week straight, believing they are millionaires and secretly married to a celebrity, etc., is not like someone having a temper tantrum.
Semantically, they can both be said to meet the criteria for schizophrenia, but these are two very different things and speaks to the limitations of the DSM. Moreover, if you take ten rare genetic variants that correlate both to schizophrenia and a mental disability, common sense should tell you that the mental disability is what is being labeled schizophrenia and not that they have both a mental disability and schizophrenia. Since schizophrenia has very specific symptoms (auditory hallucinations, paranoid delusions, etc.), it makes no sense to assume that ten different variants with entirely different neurological mechanisms all happen to lead to schizophrenia. This is not logically coherent.
What these studies are really showing, in my opinion, is one of the dirty little secrets of psychiatry and society, more generally, where if individuals are not able to cope within the acceptable structured milieu, then it falls on chemical sedation, to get them in the necessary boxes.